Future therapies in T-PLL and T-LGL

Accompanying an upcoming talk at the EHA 2021 virtual congress, our newest perspective on novel therapy approaches in mature T-cell leukemias was published in Hemasphere. In “Hijacking the Pathway: Perspectives in the Treatment of Mature T-cell Leukemias“, Linus Wahnschaffe and Marco Herling discuss future approaches for T-PLL and T-LGL that are based on current molecular disease concepts as well as on encouraging preclinical and early clinical data.
Read the perspective:
Hijacking the Pathway: Perspectives in the Treatment of Mature T-cell Leukemias
Postet on June 7, 2021New article accepted for publication
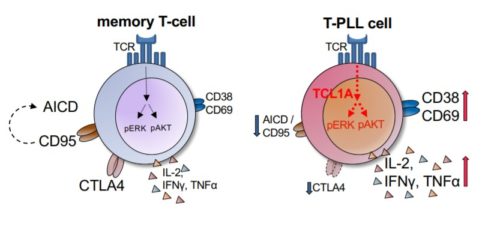
Our latest article “Non-canonical effector functions of the T-memory-like T-PLL cell are shaped by cooperative TCL1A and TCR signaling” by Sebastian Oberbeck, Alexandra Schrader, Kathrin Warner et al. has been accepted for publication in Blood.
In this study, we report two major findings:
- T-PLL cells resemble activated T-lymphocytes with augmented memory-type effector functions including a marked anergy to apoptotic triggers
- specific co-opting loss of inhibitory receptors and the overexpressed signal enhancer TCL1A lower thresholds towards permissive TCR input
We will add a link to the article as soon as the publication process has been completed.
Postet on August 30, 2020Alkylating deacetylase inhibitor tinostamustine in T-PLL
Read the article:
Postet on March 11, 2020Advances and Perspectives in the Treatment of T-PLL
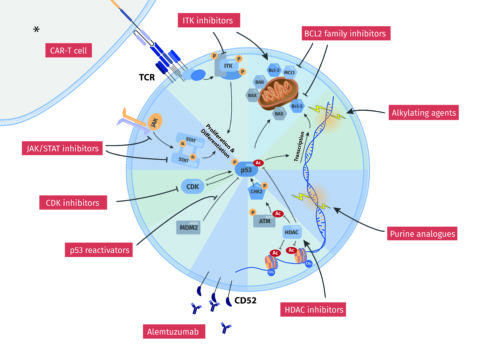
Abstract:
T cell prolymphocytic leukemia (T-PLL) is a rare mature T cell tumor. Available treatment options in this aggressive disease are largely inefficient and patient outcomes are highly dissatisfactory. Current therapeutic strategies mainly employ the CD52-antibody alemtuzumab as the most active single agent. However, sustained remissions after sole alemtuzumab-based induction are exceptions. Responses after available second-line strategies are even less durable. More profound disease control or rare curative outcomes can currently only be expected after a consolidating allogeneic hematopoietic stem cell transplantation (allo-HSCT) in best first response. However, only 30-50% of patients are eligible for this procedure. Major advances in the molecular characterization of T-PLL during recent years have stimulated translational studies on potential vulnerabilities of the T-PLL cell. We summarize here the current state of “classical” treatments and critically appraise novel (pre)clinical strategies.
Read the full article:
Advances and Perspectives in the Treatment of T-PLL
Published under a Creative Commons Attribution 4.0 International License
Postet on February 25, 2020